- 1.1 Introduction
- 1.2 Scope of Molecular Genetics
- 1.3 Deoxyribonucleic Acid (DNA)
- 1.3.1 Structure of DNA
- 1.3.2 Features of Double Helix
- 1.3.3 How DNA Decides the Hereditary Features
- 1.4 Genome
- 1.4.1 Organisation of Nuclear Genome
- 1.4.2 Organisation of Mitochondrial Genome
- 1.5 Genetic Code
- 1.5.1 Properties of Genetic Code
- 1.6 Gene Expression
- 1.6.1 Transcription
- 1.6.1.1 Ribonucleic Acid (RNA) and its Types
- 1.6.1.2 Messenger RNA (mRNA)
- 1.6.1.3 Transfer RNA (tRNA)
- 1.6.1.4 Ribosomal RNA (rRNA)
- 1.6.2 Post Transcriptional Modifications
- 1.6.3 Translation
- 1.6.3.1 Proteins
- 1.6.1 Transcription
- 1.7 Regulation of Gene Expression
1.1 INTRODUCTION
DNA is the master molecule which carries the genetic information from one generation to the other. Study of Molecular Genetics accelerated since April 25th, 1953 when James Watson and Francis Crick proposed the structure of DNA which was published in Journal called ‘Nature’. Molecular Genetics deals with the flow of genetic information and its regulation. In simple terms it can be defined as the field of biology which studies the structure and function of genes at molecular level.
1.2 SCOPE OF MOLECULAR GENETICS
Development of techniques like nucleic acid hybridisation, cloning, sequencing etc. brought a revolutionary change in Molecular Genetics. It is of major interest to the students of biology and medicine. Though it has a lot of significance in many fields, we’ll confine ourselves to the applications of molecular genetics to the mankind. They are as follows:
- i) Diagnosis of infectious diseases: Normally microorganisms are detected in the laboratory using biochemical methods. In case of molecular techniques, microorganisms are detected by using probes (short DNA or RNA sequence) which are complementary to a part of genome of the microbe. The advantage of using molecular methods is: • Identification of pathogen is done within a short time; • No need to cultivate the microbes; • Latent infections can also be identified when no antibody is formed; and • The technique can be used even when the microorganism cannot be cultured.
- ii) Diagnosis of genetic diseases : Before the advent of the above techniques, counselors used to give risk estimate like one fourth risk of getting the disease, if the parents are heterozygous for an autosomal trait. But now by directly testing for the mutation, they are able to confirm the presence or absence of mutation in the fetus. It is of immense help in prenatal diagnosis.
- iii) Individuals can be identified and relationship can be determined by DNA fingerprinting.
- iv) Mouse models for genetic diseases have been developed by creating transgenic mice.
- v) Production of vaccines, antibodies and therapeutic proteins using recombinant DNA technology. Eg; Insulin, Human Growth Hormone.
- vi)It has great potential for treating disease. Gene therapy is a process where the cells of a patient are genetically modified to alleviate disease.
- vii) With the development of recombinant DNA technology, identification of disease genes became much easier. Once the disease gene is identified, a molecular test can be designed for diagnosis of genetic disease.
| DNA Fingerprinting Just like no two individuals have identical finger prints, no two individuals have identical genetic information, except monozygotic twins. Unlike finger prints which are present only on the tips of the fingers, the genetic information which is unique to the individual is present in each and every cell. Alec Jeffrey discovered repetitive sequences called minisatellites to be unique to every individual. DNA fingerprinting is a technique which makes use of these sequences to evaluate genetic information. It’s a quick way to compare the DNA sequences of any two living organisms. It is used for personal identification, identification of the parents, when babies are switched in hospital, identification of criminals etc. |
1.3 DEOXYRIBONUCLEIC ACID (DNA)
DNA is the thread of life. It is the hereditary material in all organisms except in certain RNA viruses. All the information that is needed for the development, behavior, well being etc. of an individual is encoded in its structure. The genetic information that’s stored in DNA flows through RNA to proteins. This flow of genetic information is referred to as central dogma of molecular biology. Though the information is present in the DNA, it is the activity of proteins that is responsible for the inherited traits. The function of DNA is to direct its own replication and to direct transcription.
The number of DNA molecules present in a cell is equal to the number of chromosomes per cell. When compared to the length of the chromosome, the length of the DNA is very very long. It’s a matter of interest, how this long DNA fits into the cell whose diameter is in microns (like a long snake fitting into a small basket). This is possible because of the winding of the DNA around histone proteins into structures called nucleosomes. Nucleosomes are further coiled and coiled to form chromosome. So each chromosome is nothing but a single DNA molecule along with proteins.
1.3.1 Structure of DNA
The structure of DNA was proposed by Watson and Crick in the year 1953 for which they won Nobel Prize in 1962. The structure of DNA molecule resembles a gently twisted ladder. Two long polynucleotide chains represent the rails of the ladder. They’re coiled around a central axis to form right handed double helix. The rungs are made up of nitrogen bases which are held together by Hydrogen bonds. The basic unit of DNA is nucleotide. It’s composed of three subunitsnitrogen base, sugar and phosphate.
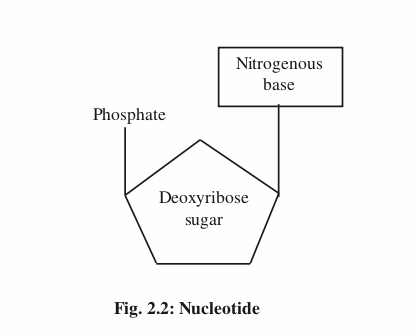
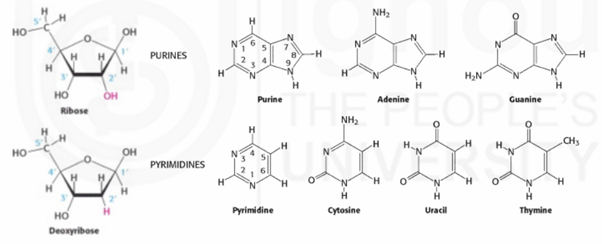
There are two kinds of bases- purines (double ringed) and pyrimidines (single ringed). A purine (Adenine and Guanine) always pairs with a pyrimidine (Thymine and Cytosine). Therefore, the amount of purines present in a DNA molecule is equal to the amount of pyrimidines. Adenine always pairs with Thymine with two Hydrogen bonds whereas Guanine always pairs with Cytosine with three Hydrogen bonds. The two strands of the DNA can be separated easily during replication because of the weak Hydrogen bonds.
The sugar is a pentose sugar which is deoxyribose in DNA because it lacks Oxygen at second Carbon position. To distinguish between the Carbon atoms present in the base and sugar, the Carbon atoms in the sugar are given a prime (‘). The bases attach to the sugar (at 1’C) by glycosidic bond.
The phosphate group links the 3’C atom of one sugar with the 5’C atom of adjacent sugar by a phospodiester bond. Because of the negative charges present on the phosphate groups, DNA is a polyanion. In vivo, these charges are neutralised by the positively charged histone proteins.
1.3.2 Features of Double Helix
The two strands of the double helix are antiparallel ie., the 5’ end of one strand aligns with 3’ end of other strand.
Because of the specific base pairing, the two strands are complementary to each other which means that if we know the sequence of one strand we can infer the sequence of the other.
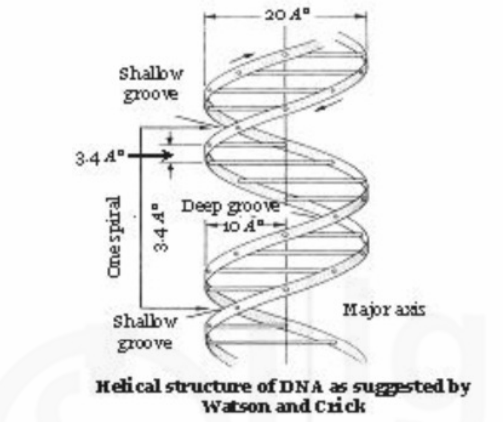
The bases are stacked on one another 3.4Ao apart and are perpendicular to the axis of the double helix. The diameter of the helix is about 20Ao and each complete turn of helix measures 34Ao, thus accommodating 10 base pairs in each turn.
Stacking of base pairs (bp) results in major and minor grooves in DNA. Major groove is rich in chemical information and is recognised by sequence specific DNA binding proteins.
1.3.3 How DNA Decides the Hereditary Features
The structure of DNA is the same in all organisms with same four nitrogenous bases-A,T,G and C. Then what makes the difference between plants and animals or how does a zygote know to develop into a monkey or a human? It’s the order of the base sequence that makes all the difference. It’s not the same in all. The bases are present in different amounts in different species.
1.4 GENOME
A genome is the total genetic information present in a cell. Basing on the complexity of humans, if you assume that among all species, the human beings have the largest amount of DNA, you’re mistaken. This is because many plant species have much more DNA per cell compared to humans. Even among vertebrates, it’s the amphibians which have the greatest amount of DNA per cell. The organisation of human genome is very complex. It comprises of two genomes (Nuclear and Mitochondrial).
1.4.1 Organisation of Nuclear Genome
The nuclear genome constitutes more than 99% of the total genome. The haploid genome contains 3 billion bp. The haploid genome is distributed in 23 different types of chromosomes (22 autosomes and 1 allosome). Each chromosome contains many genes. The genes are not uniformly distributed on the chromosomes. A certain area of the chromosome may be rich in genes while areas like centromere and telomeres are largely devoid of genes. Some chromosomes are rich in genes (22nd chromosome) some are gene poor (4th chromosome). The genes which are part of same metabolic pathway may be on different chromosomes and genes which are no way connected to metabolic pathway may be side by side on the chromosome.
There is tremendous variation in the size of the gene, size of the exon as well as intron. On an average an exon may contain < 200bp. Size of the intron may vary from 100bp to >100,000bp. About 1.5% of the total genome is coding (Exon is the coding region and Intron is non coding).
A number of protein coding genes in the human genome form gene families. A set of genes which code for similar protein sequences or which have nucleotide sequence similarity form a gene family (just like related individuals make a family). They arose by duplication of the ancestral gene and accumulation of independent mutations over a period of time. Eg: members of beta globin gene family. An individual won’t have same beta globin throughout his development. Apart from beta globin (â) there are different genes like ä, Gã, Aã and å which code for slightly different polypeptides. They express during different stages of development of an individual and forms a gene family. Members of a gene family generally appear as a cluster or they may be dispersed.
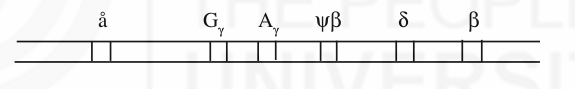
More than half of the genome contains repetitive sequences. Basing on the number of copies per genome, the DNA sequences are classified into-unique sequences (1-10 copies), moderately repetitive sequences (10-105copies) and highly repetitive sequences (>105copies). Unique sequences include most of the genes which code for proteins. Example for moderately repetitive sequences is the genes which code for ribosomal RNA and histone proteins. Highly repetitive sequences are tandemly arranged and are transcriptionally inactive. They are once again classified into mega satellite, satellite, mini satellite and micro satellite according to the decreasing size of the repeat. Mega satellites are very few in number. Satellite DNA is present in the centromeric region of the chromosomes. The length of the mini satellite DNA is quite variable among individuals and is the basis for DNA fingerprinting. Microsatellites constitute single base runs, di, tri and tetra nucleotide repeats.
Transposons, the DNA sequences which are capable of moving from one part of the genome to other constitute about 45% of the total human genome. Most of them are nonfunctional. Transposition is RNA mediated ie., DNA is first transcribed into RNA and then reverse transcribed into cDNA which is the double stranded form is inserted elsewhere in the genome.
There are pseudogenes in the genome which are nonfunctional copies of a functional gene eg. pseudo beta(øâ) in beta globin gene family. They also arose by duplication of the ancestral gene but in course of time accumulated mutations which rendered them non functional. Few overlapping genes (in class III region of HLA complex present on 6th chromosome.) and genes within genes (presence of two genes within the intron of clotting factor VIII gene) also exist in the genome.
1.4.2 Organisation of Mitochondrial Genome
The total amount of mitochondrial genome is <1%. It varies per cell basing on the number of mitochondrial DNA(mt DNA) molecules per mitochondrion and the number of mitochondria per cell.
The size of the human mtDNA is 16,569 bp. It is circular and double stranded. It lies naked in the organelle. mtDNA isn’t associated with histone proteins. It contains a light chain and a heavy chain. Heavy strand is rich in guanines and light strand is rich in cytosines. DNA is triple stranded at a region, due to duplication of a section of heavy strand and is called D loop. D loop has no coding sequences. mtDNA has altogether 37 genes out of which 13 code for polypeptides, 22 for tRNAs and 2 for rRNAs. In contrast to nuclear genome, about 93% of the mtDNA is coding. The genome is compact with no introns and presence of overlapping genes. Mitochondria have their own ribosomes on which polypeptides are synthesized. It has a slightly different genetic code when compared to that followed by the nuclear genome.
The mitochondria synthesize only some of the proteins needed by it, others being synthesized by the nuclear genes. The proteins produced by the nuclear genome are imported into mitochondria. mtDNA shows maternal inheritance because all the mitochondria received by the zygote are from the ovum. Mutations in mtDNA are responsible for certain diseases in humans.
1.5 GENETIC CODE
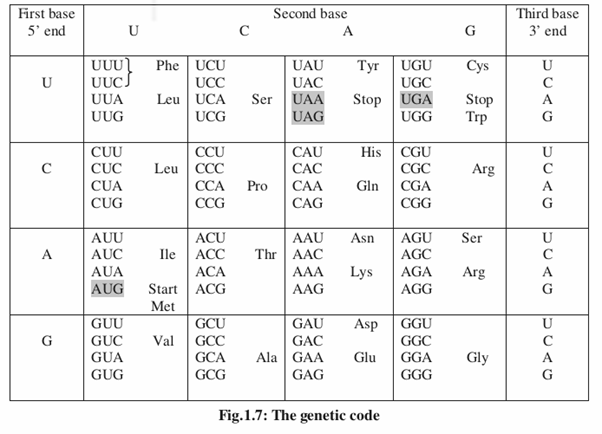
We now know that DNA contains the information that is necessary for the production of proteins. The question is how the information stored in DNA can be decoded into a protein? One of the two DNA strands is transcribed into RNA. This RNA which contains the coded information, acts as a messenger molecule which is further translated into polypeptide. It’s essential to understand the nature of genetic code to understand how the coded information in RNA is decoded to protein. Genetic code is a dictionary for the translation of mRNA into protein.
DNA is made up of only 4 different nucleotides (A,T,G and C) and proteins are synthesized from 20 different amino acids. The question is how 4 nucleotides could specify 20 amino acids ? A singlet code (each nucleotide codes for one amino acid) specifies only 4 amino acids, a doublet code (2 bases code for one amino acid) specifies only 16 amino acids (42). So the minimum number of nucleotides needed to code for 20 different amino acids is 3. This group of 3 nucleotides or nucleotide triplet is called a codon. A triplet code will contain 64 codons (43) which are in excess of the number of amino acids. The code was deciphered in 1960s by the important contributions made by Nirenberg, Matthaei, Gobind Khorana and Ochoa.
You may have a doubt whether the same genetic code is followed by plants, animals and bacteria as all of them have same 4 bases in their genetic material. Yes, genetic code is universal, except slightly different code is used in mitochondria and by few prokaryotes. Because of this property, we are able to translate mRNA from one species, in a cell of another species (recombinant DNA technology).
1.5.1 Properties of Genetic Code
The genetic code is triplet : The code is read in 3 letter words. A group of three nucleotides code for one amino acid.
The code is degenerate : There are 64 codons but amino acids are 20 only which means some amino acids are specified by more than one codon. Eg: GUU,GUC,GUA and GUG code for valine. All these 4 codons are said to be degenerate.
The code has polarity: Codons may specify different amino acids when they are read in opposite directions.
Eg.: 5’ CCU 3’ → proline
3’ UCC 5’ → serine
As translation occurs in 5’→ 3’ direction, it’s apt to read the codons in 5’→3’ direction only.
The code is non-overlapping: Each codon consists of three consecutive nucleotides. That is none of the nucleotide is part of 2 codons. Eg. 5’ GCUACCUGC 3’
Non overlapping code will specify only three amino acids. NH2-ala-thr-cys-COOH.
Overlapping code will specify seven amino acids. NH2-ala-leu-tyr-thr-pro-leucys-COOH.
The code is comma less: There is no gap or punctuation between 2 codons. After one amino acid is coded, the second one will be coded automatically. If there is any gap between two codons, deletion or addition of one base should not change the reading frame. But a change in reading frame was observed which means that code is comma less.
| Reading Frame The possible way in which a nucleotide sequence is read during translation is called the reading frame. Basing on the starting point, a single strand of DNA molecule can be read in three possible ways. Eg: 5’AGCGCAAGGCGA…..3’ The above sequence has three possible reading frames –one starting with the first base, other two frames starting with second and third bases. 5’AGC GCA AGG CGA….3’ 5’GCG CAA GGC GA….3’ 5’CGC AAG GCG A…..3’ |
The code is unambiguous: Though there are few exceptions, a particular codon will always specify the same amino acid. Eg. GCU always codes for alanine.
The code contains “start” and “stop” signals: There is only one start codon (AUG) whereas termination codons (UAA, UAG and UGA) are three in number.
Wobble Hypothesis: Leaving the three termination codons which are recognised by proteins, the remaining 61 codons are recognised by tRNAs. There are only about 30 types of cytoplasmic tRNAs. Then how’s it possible to interpret 61 codons? This is possible because of the relaxation of normal base pairing rules when it comes to codon-anticodon recognition. According to Wobble hypothesis of Crick, normal A-U and G-C rules are followed for the first two base positions only but wobbling occurs at third position (G can pair with C or U and U can pair with A or G).
1.6 GENE EXPRESSION
Before going to gene expression, let us first understand the meaning of gene, the number of genes in humans, their location and their structure. In simple terms, gene is a stretch of DNA that carries the information necessary for the synthesis of a polypeptide. Actually the definition of gene is much more complex. Today we know that a single gene can give rise to many polypeptides. There are about 25,000 genes in humans according to Human Genome Project. The number of proteins about two lakhs is far greater than the number of genes because of alternative splicing. Genes are located on chromosomes. Each chromosome contains many number of genes arranged in a linear order. Eukaryotic genes are split genes, which means that their coding sequence is not contiguous but it is interrupted by noncoding or intervening sequences called introns. The coding sequences or the expressed sequences are called as exons. In addition to the coding and noncoding sequences, there are flanking regions which are important in regulation and have ‘start’ and ‘stop’ signals. These include promoter which is located at the 5’ end of the gene and a sequence that is present at the 3’ end which provides the signal for the addition of poly A tail to the 3’ end of mature mRNA.
Gene expression is a process in which a protein is synthesized from a gene. It occurs in two major steps. The first step is transcription, in which the linear DNA is transcribed into linear mRNA. The second step is translation during which mRNA associates with the ribosomes present in the cytoplasm and directs the synthesis of proteins.
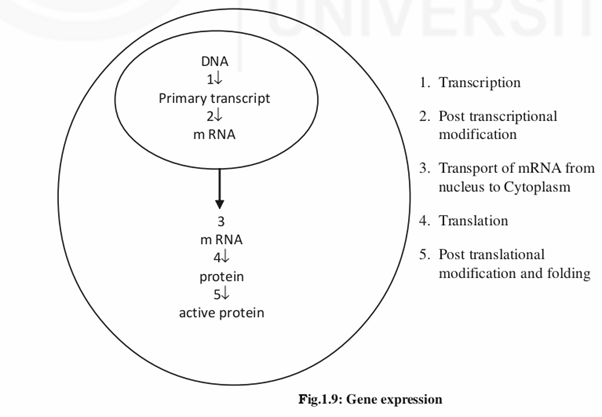
Here you should note the point that only a small proportion of the total DNA (1.5%) is coding. Moreover, all the genes that are transcribed are not translated that is the end product of some genes is RNA itself eg: tRNA, rRNA etc.
1.6.1 Transcription
It’s a process in which single stranded RNA is generated from one of the strands of the DNA. It occurs in nucleus in 5’→ 3’ direction. It needs RNA polymerase, ribonucleotides and several proteins for initiation. Only one of the two strands of the DNA acts as a template. RNA that is synthesized is complementary to the template but similar in sequence and orientation to that of nontemplate strand (except U is present in place of T). Therefore nontemplate strand is called sense strand and template strand is called antisense strand. Whenever we want to give a gene sequence, it’s customary to give the sequence of sense strand in 5’ to 3’ direction.
Nontemplate (sense strand) 5’___________________________3’
ACATGCCTATACCGACCAGCTATT DNA
Template (antisense strand) 3’TGTACGGATATGGCTGGTCGATAA 5’
5’____________________________3’ RNA
ACAUGCCUAUACCGACCAGCUAUU
The first base that’s transcribed is denoted +1 and the bases that are proceeding in the right side (5’→ 3’) are indicated by positive numbers and the direction is called downstream. Conversely the bases towards the left side of +1 are indicated by negative numbers and the direction is called upstream. The promoter which is present in the upstream region of the sense strand contains a group of short sequence elements called TATA box, GC box, CAAT box etc. These elements will be recognised by proteins called transcription factors (TFs). Only when TFs bind to the promoter, followed by binding of RNA polymerase, then transcription occurs. There are three kinds of RNA polymerases in humans. RNA polymerases I and III transcribe the genes which code for tRNA, rRNA and various small RNAs. Structural genes (genes which code for proteins) are transcribed by RNA polymerase II. Termination occurs in them by endonucleolytic cleavage (downstream to a sequence AAUAAA) followed by addition of poly(A) tail.
1.6.1.1 Ribonucleic Acid (RNA) and its Types
RNA molecule is also a polynucleotide chain. The sequence of RNA is determined by the DNA sequence. The difference between DNA and RNA is, it’s single stranded, contains ribose sugar (2’OH) in place of deoxyribose sugar (2’-H) and thymine is replaced by Uracil. Occasionally it may fold on itself to give stem loop structures.
Transcription leads to the synthesis of several different types of RNA. Messenger RNA, ribosomal RNA and transfer RNA are the major classes of RNA involved in protein synthesis.
1.6.1.2 Messenger RNA (mRNA)
It’s this RNA, which carries the message present in the gene to the cytoplasm where synthesis of protein occurs. Only the central part of the mRNA is translated. The region of the first exon and the last exon which are not translated are denoted as 5’UTR and 3’UTR. Each group of three mRNA bases constitutes a codon which specifies an amino acid. The length of different mRNAs vary considerably basing on the length of the gene.
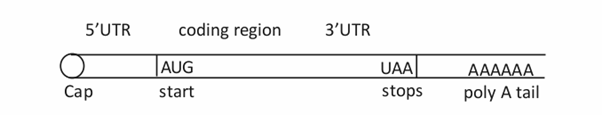
1.6.1.3 Transfer RNA (tRNA)
Transfer RNA molecules interpret the mRNAs with the aid of ribosomal RNAs. They are smallest RNAs (about 80 nucleotides long) which carry the amino acids to the site of protein synthesis. Their one end binds to a codon in the mRNA and the opposite end carries a specific amino acid. Thus it acts like an adaptor. Because of the formation of Hydrogen bonds between some of the complementary bases it forms a clover leafed structure (secondary structure). Its three dimensional structure is however L shaped.
1.6.1.4 Ribosomal RNA (rRNA)
About 80% of the total RNA is rRNA. Generally the largest of the RNAs, along with ribosomal proteins it forms the ribosome. A ribosome is made up of two subunits, both of which join at the time of protein synthesis.
Larger subunit: 3 kinds of rRNAs (28S, 5.8S and 5S) + about 50 ribosomal proteins. Smaller subunit: single rRNA (18S) + more than 30 ribosomal proteins
1.6.2 Post Transcriptional Modifications
All the above three types of RNAs undergo post transcriptional modifications that is; certain bases present in the RNAs are removed. The RNA that is obtained after transcription is termed primary transcript. In case of mRNA, the introns are removed and the exons are spliced together to form the mature mRNA. Splicing occurs with the help of certain conserved sequences present in the introns. In addition to splicing , in case of mRNA, a cap and a poly A tail are added to the 5’ and 3’ ends of the mRNA respectively. These two structures help in the migration of the mRNA from the nucleus to the cytoplasm and also in the regulation of gene expression.
We’ve already discussed about alternative splicing due to which we are getting about 2,00,000 proteins from the 25,000 genes we have. In simple terms, alternative splicing means getting more number of mRNAs from a single gene. This is possible by differential splicing in different tissues. For eg. a sequence which acts as an intron in one tissue may act as a coding sequence in another tissue thereby changing the sequence of the polypeptide in different tissues.
1.6.3 Translation
It’s a process in which the information present in an mRNA is decoded into the amino acid sequence of a protein. It requires mRNA, tRNA, ribosomes, ATP and various protein factors. It occurs on ribosomes in the cytoplasm. It also occurs in 5’→3’ direction. The 5’ end of mRNA corresponds to the amino terminus of the protein. Translation starts from the initiation codon and ends with the termination codon. That’s the reason why most of the polypeptides start with methionine.
Initiation of translation requires several factors which include a cap binding protein, initiation factors, smaller subunit of the ribosome, initiator methionyl tRNA, all of which bind to the 5’ cap region of the mRNA. The initiation complex formed scans the mRNA for the initiation codon. In the elongation step, larger subunit attaches to the initiation complex. The codon next to the AUG is then recognised by another tRNA which brings the second amino acid of the polypeptide chain. A peptide bond is formed between the two amino acids and successive amino acids are incorporated into the growing polypeptide chain. This process continues till the termination codon is reached which is recognised by a protein but not tRNA. Finally the last tRNA will be released from the ribosome the two subunits of the ribosome separate and the new polypeptide will be released.
1.6.3.1 Proteins
A protein is a polymer made up of amino acids. It’s the end product of most of the genes. Proteins perform all the metabolic reactions that are carried out in a cell. Though the term protein and polypeptide are loosely used, there is a difference between the two. Polypeptide is the molecule that’s formed after translation. After its release, the nascent polypeptide folds up and achieves a three dimensional conformation to become functional protein. Many proteins depend on other proteins called chaperones for folding. In addition to proper folding, polypeptides also undergo post translational modifications (hydroxylation, glycosylation, phosphorylation etc.) to achieve functional status. So a polypeptide is a precursor of protein. Some proteins may have more than one polypeptide, which may be of same kind or of different kinds.
1.7 REGULATION OF GENE EXPRESSION
There are 200 different types of cells in human beings. All of them have the same DNA content or to be precise all the genes. Yet all of them are morphologically different and have different function. This depends on the type of genes that are expressed in these cells or in other words differential gene expression is responsible for the diverse properties of different cells. All the genes are not expressed in all the cells or all the times. By this we mean that some genes are expressed only in some cells (tissue specific expression). Similarly some genes are expressed only during a particular time of development. However there are certain genes which are expressed in all cell types (house keeping genes) eg: genes for rRNA, tRNA, DNA polymerases etc. The question is how does a cell know which genes to express, when to express and to what extent. That’s what is explained in this topic.
It’s a waste of energy for the cell to produce the proteins which are not needed by it. Cells have their own methods, by which they can regulate the expression of genes. Interestingly it’s the proteins which are largely responsible for regulation of gene expression. Regulation occurs at three levels.
1) Transcription is the predominant stage at which regulation occurs. Transcription occurs at basal level with the help of TFs. Up regulation and down regulation of gene expression is possible with the help of proteins called activators and repressors respectively. Activators bind to sequences called enhancers whereas repressors bind to silencers. These sequences may be located near the promoter region or far away in the upstream or downstream region of the gene. These regulatory proteins are controlled by signals which determine whether these proteins bind DNA. They determine the amount of the protein to be synthesized. Tissue specific expression is possible by limiting the availability of TFs needed by a gene only to a particular tissue. In some cases, the promoter sequence is methylated in all other tissues except the tissue where it’s expressed. Histone proteins also have a role in regulation. Methylation of certain amino acids in histone proteins turns off the expression of a gene.
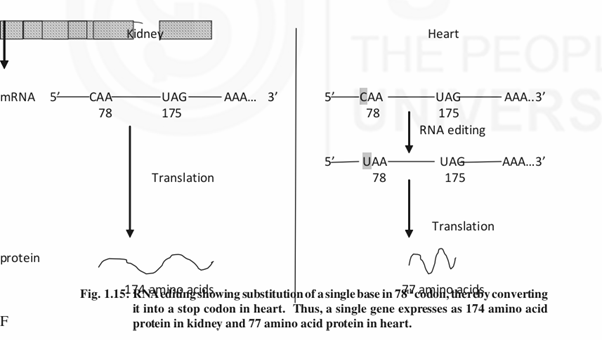
2) Regulation also occurs at post transcriptional level. Alternative splicing produces different isoforms in different tissues. Isoforms also result due to alternative polyadenylation (same gene uses different polyadenylation signals in different tissues) and RNA editing (same gene produces different isoforms due to single base substitution, deletion or insertion at the RNA level) .
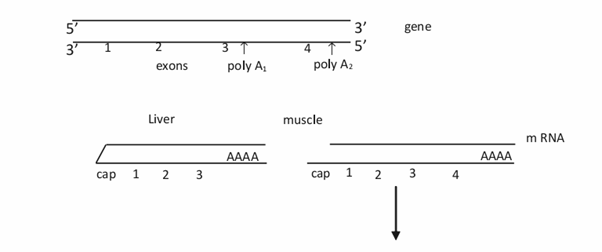
3) Regulation at translation level includes longevity of mRNA which depends on length of poly A tail (mRNAs without poly A tail are short lived), structure of 3’UTR (many repeats of AUUUA in the 3’UTR makes the RNA short lived) etc. Translation of some mRNAs is regulated by specific RNA binding proteins. Degradation of mRNA is another control point. Rapid degradation of mRNA prevents undesired protein synthesis.
Sample Questions
- 1) Describe the salient features of DNA double helix.
- 2) “The organisation of human nuclear genome is complex”. Justify the statement.
- 3) Define genetic code and write about its properties.
- 4) Explain how the nucleotide sequence present in a gene is used to synthesize a protein.
- 5) Describe how gene expression is regulated at different levels.
Short Notes
- i) MtDNA
- ii) Genome
- iii) RNA and its Types