As we defined it in Chapter 4, a population is a group of interbreeding individuals. More precisely, a population is the group within which an individual is most likely to find a mate. As such, a population is marked by a degree of genetic relatedness and shares a common gene pool.
In theory, this is a straightforward concept. In every generation, the genes (alleles) are mixed by recombination and rejoined through mating. What emerges in the next generation is a direct product of the genes going into the pool, which in turn is a product of who is mating with whom.
In practice, however, describing human populations is difficult. The largest human population that can be described is our entire species. All members of a species are potentially capable of interbreeding but are incapable of producing fertile offspring with members of other species. Our species, like any other, is thus a genetically closed system. The problem arises not in describing who can potentially mate with whom but in determining the exact pattern of those individuals who are doing so.
Factors that determine mate choice are geographical, ecological, and social. If individuals are isolated on a remote island in the middle of the Pacific Ocean, there isn’t much chance that they’ll find mates outside the immediate vicinity. Such breeding isolates are fairly easily defined and are a favorite focus of microevolutionary studies. Geography plays a dominant role in producing these isolates by severely limiting the range of available mates. But even within these limits, cultur al rules can easily play a deciding role by stipulating who is most appropriate among those who are potentially available.
Human population segments are defined as groups with relative degrees of endogamy (marrying/mating within the group). But these aren’t totally closed systems. Gene flow often occurs between groups, and individuals may choose mates from distant locations. With the advent of modern transportation, the rate of exogamy (marrying/ mating outside the group) has dramatically increased.
Today, most humans aren’t clearly defined as members of particular populations because they don’t belong to a breeding isolate. Inhabitants of large cities may appear to be members of a single population; but within the city, there’s a complex system of social, ethnic, and religious boundaries that are crosscut to form smaller population segments. Besides being members of these highly open local population groupings, we’re simultaneously members of overlapping gradations of larger populations—the immediate geographical region (a metropolitan area or perhaps an entire state), a section of the country, the entire nation, and ultimately the whole species.
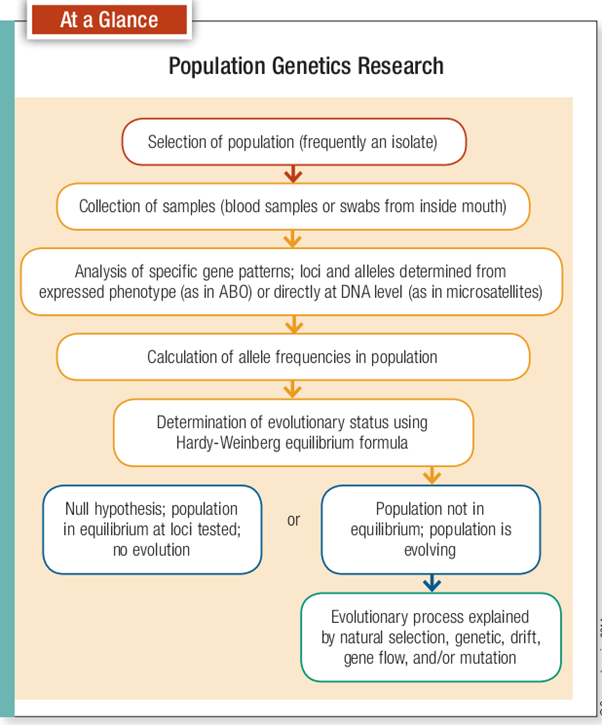
After identifying specific human populations, the next step is to find out what evolutionary forces, if any, are operating on them. To determine whether evolution is taking place at a given genetic locus, we measure allele frequencies for specific traits. We then compare these observed frequencies with those predicted by a mathematical model called the Hardy-Weinberg theory of genetice quilibrium. This model gives us a baseline set of evolutionary expectations under known conditions.
The Hardy-Weinberg theory establishes a set of conditions in a hypothetical population where no evolution occurs. In other words, no evolutionary forces are acting and all genes have an equal chance of recombining in each generation (that is, there’s random mating of individuals). More precisely the conditions that such a population would be assumed to meet are as follows:
- 1. The population is infinitely large; this eliminates the possibility of random genetic drift—that is, changes in allele frequencies due to chance.
- 2. There’s no mutation; thus, no new alleles are being added by changes in genes.
- 3. There’s no gene flow; thus there’s no exchange of genes with other populations that could alter allele frequencies
- 4. Natural selection isn’t operating; thus specific alleles offer no advantage over others that might influence reproductive success.
- 5. Mating is random; therefore, there’s nothing to influence who mates with whom; all females are assumed to have an equal chance of mating with any male, and vice versa.
If all these conditions are met, allele frequencies won’t change from one generation to the next (that is, no evolution will take place), and as long as these conditions prevail, the population maintains a permanent equilibrium. This equilibrium model provides population geneticists with a standard against which they can compare actual circumstances. Notice that the conditions defining the Hardy-Weinberg equilibrium constitute an idealized, hypothetical state. In the real world, no actual population would fully meet any of these conditions. But don’t be confused by this distinction. By explicitly defining the allele frequencies that would be expected if no evolutionary change were occurring (that is, in equilibrium), we establish a baseline with which to compare the allele frequencies we actually observe in real human populations.
If the observed frequencies differ from those of the expected model, we can then say that evolution is taking place at the locus in question. The alternative, of course, is that the observed and expected frequencies don’t differ enough that we can confidently say that evolution is occurring at a particular locus in a population. In fact, this is often what happens; in such cases, population geneticists aren’t able to clearly define evolutionary change at the particular locus under study.
The simplest way to do a micro evolutionary study is to observe a genetic trait that follows a simple Mendelian pattern and has only two alleles (A and a). Remember that there are only three possible genotypes: AA, Aa, and aa. Proportions of these genotypes (AA:Aa:aa) are a function of the allele frequencies themselves (percentage of A and percentage of a). To provide uniformity for all genetic loci, a standard notation is employed to refer to these frequencies:
Frequency of dominant allele (A) = p
Frequency of recessive allele (a) = q
Since in this case there are only two alleles, their combined total frequency must represent all possibilities. In other words, the sum of their separate frequencies must be 1:
p + q
(Frequency of A alleles) (Frequency of = A alleles)
1 (100% of alleles at the locus in question)
To determine the expected proportions of genotypes, we compute the chances of the alleles combining with one another in all possible combinations. Remember, they all have an equal chance of combining and no new alleles are being added. These probabilities are a direct function of the frequency of the two alleles. The chances of all possible combinations occurring randomly can be simply shown as
(p + q) × ( p + q ) = (pq + q²) +( p² + pq) = P² + 2pq + q²
Mathematically, this is known as a binomial expansion and can also be shown as:
(p + q)(p + q) = p² + 2pq + q²
What we have just calculated is simply: Expected
Allele Combination Genotype Produced Proportion in Population
Chances of AA p × p = p² A combining with A
Chances of Aa p × q = A combining with a; 2pq a combining aA p × q = with A Chances of aa q × q = q² a combining with a
Thus, p² is the frequency of the AA genotype, 2pq is the frequency of the Aa genotype, and q² is the frequency of the aa genotype, where p is the frequency of the dominant allele and q is the frequency of the recessive allele in a population
Calculating Allele Frequencies
We can best demonstrate how geneticists use the Hardy-Weinberg formula by giving an example. Let’s assume that a population contains 200 individuals, and we’ll use the MN blood group locus as the gene to be measured. The two alleles of the MN locus produce two antigens (M and N) that are similar to the ABO antigens and are also located on red blood cells. Because the M and N alleles are codominant, we can ascertain everyone’s phenotype by taking blood samples and testing them in a process very similar to that for ABO (see Fig. 14-2). From the phenotypes, we can then directly calculate the observed allele frequencies. So let’s see what we can determine.
All 200 individuals are tested, and the observed data for the three phenotypes are as follows:
Genotype Number of individuals* Percent number of alleles
Individual
MM 80 40 160 0
MN 80 40 80 80
NN 40 20 0 80
Totals 200 100 240 + 160 = 400
Proportion 0.6 + 0.4 = 1
From these observed results, we can count the number of M and N alleles
and thus calculate the observed allele frequencies:
p = frequency of M = 0.6
q = frequency of N = 0.4
The total frequency of the two alleles combined should always equal 1. As you can see, they do.
Next, we need to calculate the expected genotypic proportions. This calculation comes directly from the Hardy-Weinberg equilibrium formula:
p² + 2 pq + q² = 1.
p² = (.6) (.6) = .36
2 pq = 2(.6) (.4) = 2(.24) = .48
q² = (.4)(.4) = .16
Total 1.00
There are only three possible genotypes: MM, MN, and NN. The total of the relative proportions should equal 1. Again, as you can see, they do. Finally, we need to compare the two sets of data—that is, the observed frequencies (what we actually found in the population) with the expected frequencies (those predicted by HardyWeinberg under conditions of genetic equilibrium). How do these two sets of data compare?
Expected Number of observed Actual Number
Frequency Individuals Frequency of Individuals with
Each Genotype MM 36 72 40 80
MN 48 96 40 80
NN 16 32 20 40
We can see that although the match between observed and expected frequencies isn’t perfect, it’s close enough statistically to satisfy equilibrium conditions. Since our population isn’t a large one, sampling may easily account for the small degree of observed differences. Our population is therefore probably in equilibrium (that is, it’s not evolving at this locus). Of course, the observed allele frequencies do sometimes vary enough from equilibrium predictions to suggest that the population isn’t in equilibrium—that is, it’s evolving. For example, consider the locus influencing PTC tasting. What makes PTC tasting such a useful characteristic is how easy it is to identify. Unlike blood antigens such as ABO or MN, PTC tasting can be tested by simply having subjects place a thin paper strip on their tongues. This paper contains concentrated PTC, and people either taste it or they don’t.
So testing is quick, inexpensive, and doesn’t involve a blood test. With such an efficient means of screening subjects, we now consider a sample population of 500 individuals. The results from observing the phenotypes and calculations of expected genotypic proportions are shown in “A Closer Look: Calculating Allele Frequencies: PTC Tasting in a Hypothetical Population.” You’ll find additional examples of population genetics calculations in Appendix D, which is available on the Anthropology CourseMate at Cengagebrain.com.
Evolution in Action: Modern Human Populations
Once a population has been defined, it’s possible to determine whether allele frequencies are stable (that is, in genetic equilibrium) or changing. As we’ve seen, the Hardy-Weinberg formula provides the tool to establish whether allele frequencies are indeed changing. But what factors cause changes in allele frequencies? There are a number of factors, including:
1. Production of new variation (that is, mutation)
2. Redistribution of variation through gene flow or genetic drift
3. Selection of “advantageous” allele combinations that promote reproductive success (that is, natural selection) Notice that factors 1 and 2 constitute the first stage of the evolutionary process, as first emphasized by the Modern Synthesis, while factor 3 is the second stage (see Chapter 4). There’s also another factor, as implied by the condition of genetic equilibrium that under idealized conditions all matings are random. Thus, an evolutionary alteration (that is, deviation from equilibrium) is called non-random mating.
Nonrandom Mating
Although sexual recombination doesn’t itself alter allele frequencies, any consistent bias in mating patterns can change the genotypic proportions. By affecting genotype frequencies, nonrandom mating causes deviations from Hardy-Weinberg expectations of the proportions p², 2pq, and q². It therefore sets the stage for the action of other evolutionary factors, particularly natural selection. A form of nonrandom mating, called assortative mating, occurs when individuals of either similar phenotypes (positive assortative mating) or dissimilar phenotypes (negative assortative mating) mate more often than expected by Hardy-Weinberg predictions. However, in the vast majority of human populations, neither factor appears to have much influence. Inbreeding is a second type of nonrandom mating, and it can have important medical and evolutionary consequences. Inbreeding occurs when relatives mate more often than expected. Such matings will increase homozygosity, since relatives who share close ancestors will probably also share more alleles than two unrelated people would. When relatives mate, their offspring have an increased probability of inheriting two copies of potentially harmful recessive alleles from a relative (perhaps a grandparent) they share in common. Many potentially deleterious genes that are normally “masked” in heterozygous carriers may be expressed in homozygous offspring of inbred matings and therefore “exposed” to the action of natural selection. Among offspring of firstcousin matings in the United States, the risk of congenital disorders is 2.3 times greater than it is for the overall population. Matings between especially close relatives (incest) often lead to multiple genetic defects. All societies have incest taboos that ban matings between close relatives, such as between parent and child or brother and sister. Thus these matings usually occur less frequently than predicted under random mating conditions. Whether biological factors also interact to inhibit such behavior has long been a topic of debate among anthropologists. For many social, economic, and ecological reasons, exogamy is an advantageous strategy for hunting and gathering bands. Selective pressures may also play a part, since highly inbred offspring have a greater chance of expressing a recessive genetic disorder and thereby lowering their reproductive fitness. What’s more, inbreeding reduces genetic variability among offspring, potentially reducing reproductive success (Murray, 1980). In this regard, it’s interesting to note that incest avoidance is widespread among vertebrates. Detailed studies of freeranging chimpanzees indicate that they usually avoid incestuous matings within their family groups, although exceptions do occur (Constable et al., 2001). In fact, in most primate species, adults of one sex consistently find mates from groups other than the one in which they were reared (see Chapter 7). As we’ve seen, recognition of close kin apparently is an ability displayed by several (perhaps all) primates. Primatologists are currently investigating this aspect of our primate cousins. Apparently both biological factors (in common with other primates) and uniquely human cultural factors have interacted during hominin evolution to produce this universal behavioral pattern among contemporary societies
Human Biocultural Evolution
We’ve defined culture as the human strategy of adaptation. Human beings live in cultural environments that are continually modified by human activity; thus evolutionary processes are understandable only within this cultural context. We’ve discussed at length how natural selection operates within specific environmental settings. For humans and many of our hominin ancestors, this means an environment dominated by culture. For example, the sickle-cell allele hasn’t always been an important genetic factor in human populations. Before the development of agriculture, humans rarely, if ever, lived close to mosquito-breeding areas. With the spread in Africa of slash-and-burn agriculture, perhaps in just the last 2,000 years, penetration and clearing of tropical rain forests occurred. This deforestation created open, stagnant pools that provided prime mosquitobreeding areas in close proximity to human settlements. DNA analyses have further confirmed such a recent origin and spread of the sickle-cell allele in West Africa. A recent study of a population from Senegal has estimated the origin of the HbS mutation in this group at between 2,100 and 1,250 ya (Currat et al., 2002). So quite recently, and for the first time, malaria struck human populations with its full impact; and it became a powerful selective force. No doubt, humans attempted to adjust culturally to these circumstances, and many biological adaptations also probably came into play. The sickle-cell trait is one of these biological adaptations. But there’s a definite cost involved with such an adaptation. Carriers have increased resistance to malaria and presumably higher reproductive success, though some of their offspring may be lost to the genetic disease sickle-cell anemia. So there’s a counterbalancing of selective forces with an advantage for carriers only in malarial environments. (The genetic patterns of recessive traits such as sickle-cell anemia are discussed in Chapter 4.) Following World War II, extensive DDT spraying by the World Health Organization began systematic control of mosquito-breeding areas in the tropics. Forty years of DDT spraying killed millions of mosquitoes; but at the same time, natural selection acted to produce several strains of DDT-resistant mosquitoes (Fig. 14-4). Accordingly, espe cially in the tropics, malaria is again on the rise, with up to 500 million new cases reported annually and more than 1 million people dying each year.
A genetic characteristic (such as sickle-cell trait) that provides a reproductive advantage to heterozygotes in certain environments is a clear example of natural selection in action among human populations. The precise evolutionary mechanism in the sickle-cell example is called a balanced polymorphism. A polymorphism, as we’ve defined it, is a trait with more than one allele at a locus in a population. But when a harmful allele (such as the sicklecell allele) has a higher frequency than can be accounted for by mutation alone, a more detailed evolutionary explanation is required. In this case, the additional mechanism is natural selection. This brings us to the second part of the term. By balanced we mean the interaction of selective pressures operating on specific alleles in a particular environment (in this case, the sickle-cell alleles in malarial areas). Some individuals (mainly homozygous normals) will die of the infectious disease malaria. Others (homozygous recessives) will die of the inherited disease sickle-cell anemia. Thus, the individuals with the highest reproductive success are the heterozygotes who have sickle-cell trait. These heterozygotes pass both the normal allele (HbA) and the sickle-cell allele (HbS) to offspring, thus maintaining both alleles at fairly high frequencies. Since one allele in this population won’t significantly increase in frequency over the other allele, this situation will become “balanced” and will persist as long as malaria continues to be a selective factor.
Lactose intolerance, which involves an individual’s ability to digest milk, is another example of human biocultural evolution. In all human populations, infants and young children are able to digest milk, an obvious necessity for any young mammal. One ingredient of milk is lactose, a sugar that’s broken down by the enzyme lactase. In most mammals, including many humans, the gene that codes for lactase production “switches off” in adolescence. Once this happens, if a person drinks fresh milk, the lactose ferments in the large intestine, leading to diarrhea and severe gastrointestinal upset. So, as you might expect, adults stop drinking fresh milk. Among many African and Asian populations (a majority of humankind today), most adults are lactose-intolerant (Table 14.1). But in other populations, including some Africans and Europeans, adults continue to produce lactase and are able to digest fresh milk. This continued production of lactase is called lactase persistence.
Evidence has suggested a simple dominant mode of inheritance for lactase persistence in adults. The environment also plays a role in expression of the trait—that is, whether a person will be lactose-intolerant— since intestinal bacteria can somewhat buffer the adverse effects of drinking fresh milk. Because these bacteria increase with previous exposure, some tolerance can be acquired, even in individuals who genetically are not lactase-persistent.
Throughout most of hominin evolution, milk was unavailable after weaning; therefore, there may be a selective advantage to switching off the gene that codes for lactase production. So why can some adults (the majority in some populations) tolerate milk? The distribution of lactose-tolerant populations may provide an answer to this question, and it suggests a powerful cultural influence on this trait. Europeans, who are generally lactose-tolerant, are partly descended from Middle Eastern populations. Often economically dependent on pastoralism, these groups raised cows and/or goats and probably drank considerable quantities of milk. In such a cultural environment, strong selection pressures apparently favored lactose tolerance, a trait that has been retained in modern Europeans. Genetic evidence from north-central Europe supports this interpretation. DNA analysis of both cattle and humans suggests that these species have, to some extent, influenced each other genetically. The interaction between humans and cattle resulted in cattle that produce high-quality milk and humans with the ability to digest it (Beja-Pereira et al., 2003). In other words, more than 5,000 ya, populations of north- central Europe were selectively breeding cattle for higher milk yields. And as these populations were increasing their dependence on fresh milk, they were inadvertently selecting for the gene that produces lactase persistence.
Table 14.1 Frequencies of Lactase Persistence
Population Group Percent
U.S. whites 81–98
Swedes and Danes >90
Swiss 88
U.S. blacks 23–30
Ibos 1
Bantu 10
Fulani 50
Chinese 1
Thais 1
Asian Americans <5
Native Americans 85
But perhaps even more informative is the distribution of lactose tolerance in Africa, where the majority of people are lactose-intolerant. Groups such as the Fulani and Tutsi have been pastoralists for perhaps thousands of years and have much higher rates of lactase persistence than nonpastoralists (Fig. 14-5). Presumably, like their European counterparts, they’ve retained the ability to produce lactase because of their continued consumption of fresh milk (Powell and Tishkoff, 2003).
Recent molecular evidence has supported this hypothesis, showing a similar coevolution of humans and cattle in East Africa (Tishkoff et al., 2007). The pattern of DNA mutations (SNPs) in Africa is different from that seen in Europe, strongly suggesting that lactase persistence has evolved independently in the two regions. In fact, the data show that lactase persistence has evolved several times just in East Africa. The importance of cattle domestication in providing milk for human groups was clearly a cultural and dietary shift of major importance. As humans selectively bred cattle to produce more and higher-quality milk, they promoted fairly rapid evolution in these animals. At the same time, humans in different areas coevolved through natural selection as allele frequencies shifted to produce higher frequencies of lactase persistence. As we’ve seen, the geographical distribution of lactase persistence is related to a history of cultural dependence on fresh milk products. There are, however, some populations that rely on dairying but don’t have high rates of lactase persistence (Fig. 14-6). It’s been suggested that such groups have traditionally consumed their milk in the form of cheese and yogurt, in which the lactose has been broken down by bacterial action.
The interaction of human cultural environments and changes in lactose tolerance in human populations is another example of biocultural evolution. In the last few thousand years, cultural factors have initiated specific evolutionary changes in human groups. Such cultural factors have probably influenced the course of human evolution for at least 3 million years, and today they are of paramount importance.