What Is a Gene?
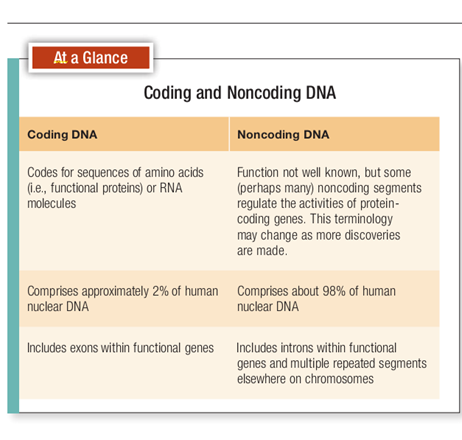
For 50 years or so, biologists considered a gene to be an uninterrupted sequence of DNA bases responsible for the manufacture of a protein or part of a protein. Or, put another way, a gene could be defined as a segment of DNA that specifies the sequence of amino acids in a particular protein. This definition, based on the concept of a one gene‒one protein relationship, was a core principle in biology for decades, but it’s been substantially modified, partly in recognition of the fact that DNA codes not only for proteins but also for RNA and other DNA nucleotides.
Moreover, when the human genome was sequenced in 2001, scientists determined that humans have only about 25,000 genes (International Human Genome Sequencing Consortium, 2001; Venter et al., 2001). This number has now been revised to approximately 21, 000 (Pennisi, 2012). Yet we produce as many as 90,000 proteins! Furthermore, protein-coding genes (also called coding sequences), the DNA segments that are transcribed into proteins, make up only about 2 to 3 percent of the entire human genome! The rest is composed of noncoding DNA or what used to be called “junk DNA” . Thus gene action is much more complicated than previously believed and it’s impossible for every protein to be coded for by a specific gene.
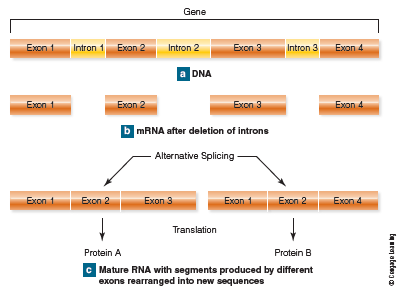
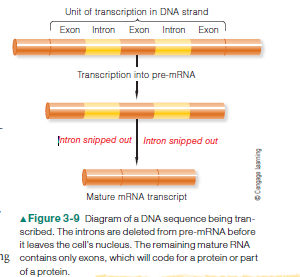
Geneticists have also learned that only some parts of genes, called exons, are actually transcribed into mRNA and thus code for specific amino acids. In fact, most of the nucleotide sequences in genes are not expressed during protein synthesis. (By expressed we mean that the DNA sequence is actually making a product.) Many sequences, called introns, are initially transcribed into mRNA and then clipped out (Fig. 3-9). Therefore introns aren’t translated into amino acid sequences. Moreover, the intron segments that are snipped out of a gene aren’t always the same ones. This means that the exons can be combined in different ways to make segments that code for more than one protein. That’s how 21,000 coding sequences can make 90,000 proteins. Genes can also overlap one another, and there can be genes within genes . But they’re still a part of the DNA molecule, and it’s the combination of introns and exons, interspersed along a DNA strand that makes up the unit we call a gene. So much for beads on a string. Clearly, the answer to the question “What is a gene?” is complicated, and a completely accurate definition may be a long time coming. However, a proposed and more inclusive definition simply states that a gene is “a complete chromosomal segment responsible for making a functional product” (Snyder and Gerstein, 2003). In spite of all the recently obtained information that has changed some of our views and expanded our knowledge of DNA, there is one fact that doesn’t change. The genetic code is universal, and at least on earth, DNA is the molecule that governs the expression, inheritance, and evolution of biological traits in all forms of life. The DNA of all organisms, from bacteria to oak trees to fruit flies to human beings, is composed of the same molecules using the same kinds of instructions. The DNA triplet CGA, for example, specifies the amino acid alanine regardless of species. These similarities imply biological relationships between all forms of life—and a common ancestry as well. What make fruit flies distinct from humans isn’t differences in the DNA material itself, but differences in how that material is arranged and regulated.
Regulatory Genes
Some genes act solely to control the expression of other genes. Basically these regulatory genes make various kinds of RNA, proteins, and other molecules that switch other DNA segments (genes) on or off. Also, many regulatory genes diminish or enhance the expression of other genes. They play a fundamental role in embryological development, cellular function, and evolution. In fact, without them, life as we know it could not exist. The study of regulatory genes and their role in evolution is still in its infancy; but as information about them continues to accumulate, we will eventually be able to answer many of the questions we still have about the evolution of species.
DNA deactivation during embryonic development is one good example of how regulatory genes work. As you know, all somatic cells contain the same genetic information; but in any given cell, only a fraction of the DNA is actually involved in protein synthesis. For example, like the cells of the stomach lining, bone cells have DNA that codes for the production of digestive enzymes. But bone cells don’t produce digestive enzymes. Instead, they make collagen, the main organic component of bone. This is because cells become specialized during embryonic development to perform only certain functions, and most of their DNA is permanently switched off by regulatory genes. In other words, they become specific types of cells, such as bone cells.
There are thousands of kinds of regulatory genes and one crucially important group is referred to as homeobox genes. The best known homeo box genes are the Hox genes, which direct the early segmentation of embryonic tissues. They also determine the identity of individual segments, by specifying what they will become, such as part of the head or thorax. Hox genes interact with other genes to determine the characteristics of developing body segments and structures but not their actual development. For example, they determine where, in a developing embryo, limb buds will appear; and they establish the number and overall pattern of the different types of vertebrae, the bones that make up the spine
All homeobox genes are highly conserved, meaning that they’ve been maintained throughout much of evolutionary history. They’re present in all invertebrates (such as worms and insects) and vertebrates, and they don’t vary greatly from species to species. This type of conservation means not only that these genes are vitally important but also that they evolved from genes that were present in some of the earliest forms of life. Moreover, changes in the behaviour of homeobox genes are responsible for various physical differences between closely related species or different breeds of domesticated animals. For these reasons, homeobox genes, and the many other kinds of regulatory genes, are now a critical area of research in evolutionary and developmental biology.
The finches of the Galápagos Islands provide an excellent example of how regulatory genes influence evolutionary change. We saw how Charles Darwin came to recognize that variation in these finches was an example of natural selection. Scientists have now explained the genetic basis for some of the finch variation by identifying two of the regulatory genes involved in the shape and size of bird beaks (Abzhanov et al., 2004, 2006). One of these genes (also involved in bone formation) is expressed to a greater degree during the embryonic development of wide beaked ground fiches than in that of finches with narrower beaks. Likewise, another gene is more active during beak development in finches that have longer, narrower beaks. Therefore the length and width of bird beaks are controlled by the activity of at least two different regulatory genes, allowing each aspect of beak size to evolve separately.
There are many other types of highly conserved genes as well. For example, recent sequencing of the sea sponge genome has shown that humans share many genes with sea sponges (Srivastava et al., 2010). This doesn’t mean that sponges were ancestral to humans, but it does mean that we have genes that were already in existence some 600 mya. These genes ultimately laid the foundation for the evolution of complex animals, and they’re crucial to many of the basic cellular processes that are fundamental to life today. These processes include a cell’s ability to recognize foreign cells (immunity), the development of specific cell types, and signalling between cells during growth and development.
We cannot overstate the importance of regulatory genes in evolution. The fact that these genes, with little modification, are present in all complex (as well as in some not so complex) organisms, including humans, is the basis of biological continuity between species.
Noncoding DNA— Not Junk After All
In all fields of inquiry, important discoveries always raise new questions that eventually lead to further revelations. There’s probably no statement that could be more appropriately applied to the field of genetics. For example, in 1977, geneticists recognized that during protein synthesis, the initially formed mrNa molecule contains many more nucleotides than are represented in the subsequently produced protein. this finding led to the discovery of introns, portions of genes that don’t code for proteins. In the 1980s, geneticists learned that only about 2 percent of human DNa is contained within exons, the segments that actually provide the code for protein synthesis. We also know that a human gene can specify the production of as many as three different proteins by using different combinations of the exons interspersed within it (pennisi, 2005).
As discussed earlier, with only 2 percent of the human genome directing protein synthesis, humans have more no protein coding DNa than any other species so far studied. Invertebrates and some vertebrates have only small amounts of noncoding sequences, and yet they’re fully functional organisms. So just what does all this noncoding DNa (originally called “junk DNa”) do in humans? Apparently much of it codes for different forms of rNa that act to regulate gene function, but it does not directly participate in protein synthesis (pennisi, 2012; the eNcODe project consortium, 2012).
Almost half of all human DNa consists of noncoding segments that are repeated over and over and over. Depending on their length, these segments have been referred to as tandem repeats, satellites, or microsatellites, but now they’re frequently lumped together and called copy number variants (cNVs). Microsatellites have an extremely high mutation rate and can gain or lose repeated segments and then return to their former length. But this tendency to mutate means that the number of repeats in a given microsatellite varies between individuals. and this tremendous variation has been the basis for DNa fingerprinting, a technique commonly used to provide evidence in criminal cases. actually, anthropologists are now using microsatellite variation for all kinds of research, from tracing migrations of populations to paternity testing in nonhuman primates. Some of the variations in microsatellite composition are associated with various disorders, so we can’t help wondering why these variations exist. One answer is that some microsatellites influence the activities of protein coding DNa sequences. also, by losing or adding material, they can alterthe sequences of bases in genes, thus becoming a source of mutation in functional genes. and these mutations are a source of genetic variation.
Lastly, there are transposable elements (tes), the so-called jumping genes. these are DNa sequences that can make thousands of copies of themselves, which are then scattered throughout the genome. One family of tes, called alu, is found only in primates. about 5 percent of the human genome is made up of alu sequences, and although most of these are shared with other primates, about 7,000 are unique to humans (chimpanzee Sequencing and analysis consortium, 2005).
TEs mainly code for proteins that enable them to move about, and because they can land right in the middle of coding sequences (exons), tes cause mutations. Some of these mutations are harmful, and tes have been associated with numerous disease conditions, including some forms of cancer (Deragon and capy, 2000). But at the same time, tes essentially create new exons, thereby generating variations on which natural selection can act. Moreover, they also regulate the activities of many genes, including those involved in development. So rather than being junk, are increasingly being recognized as serving extremely important functions in the evolutionary process, including the introduction of genetic changes that have led to the origin of new lineages.