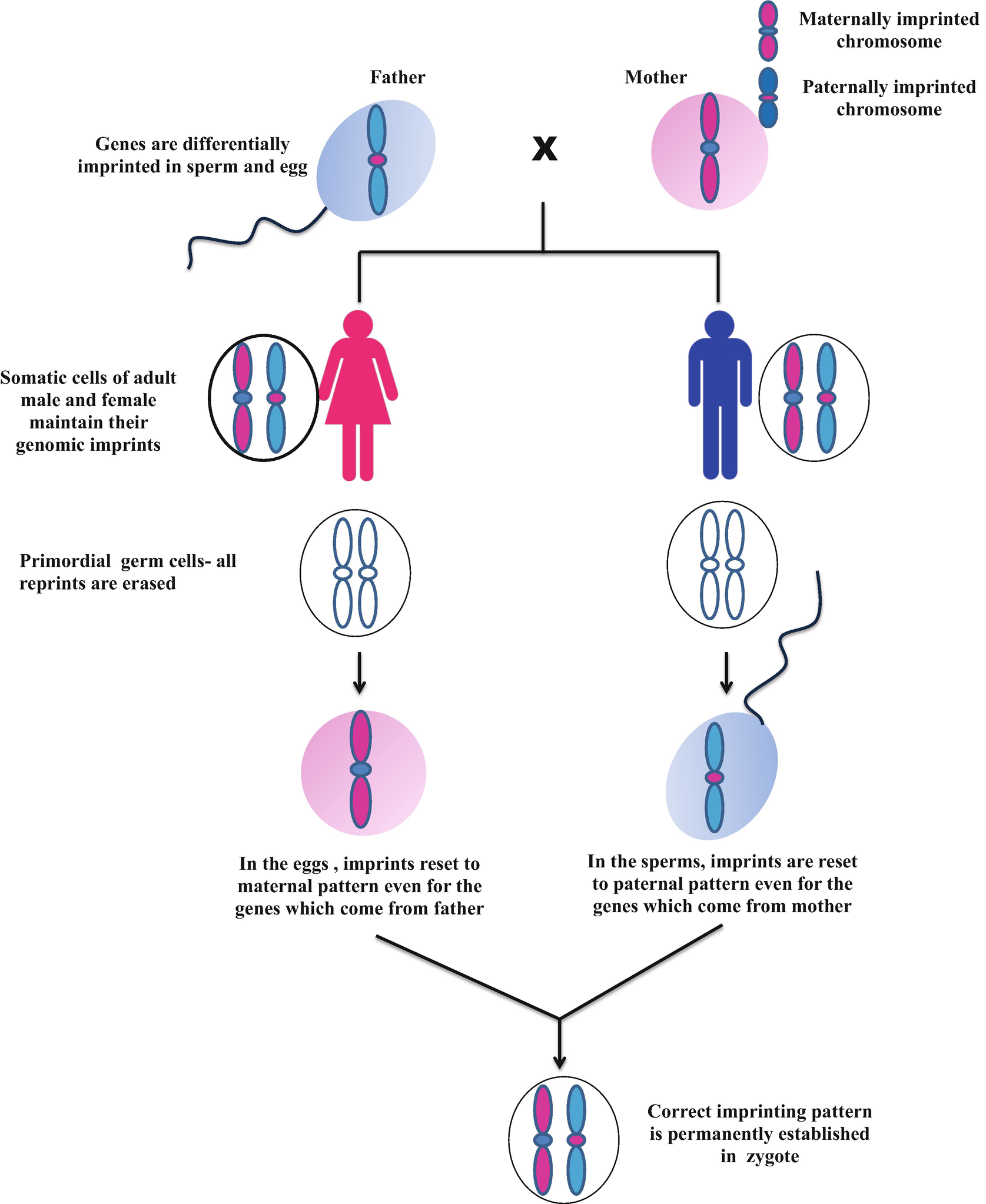
Genomic imprinting refers to a process whereby the maternal copy of a gene can be marked or “imprinted” differently than the paternal copy of the same gene (Reik and Walter, 2001). The imprinting process marks a relatively small number (100–200) of human genes and occurs in the germline. During the early stages of oogenesis, the paternal mark/imprint on a gene (coming from the father’s chromosome) is erased and a maternal imprint is then placed on the gene. In this way, both gene copies in the egg are properly marked as maternal. Conversely, in spermatogenesis, the maternal imprint of a gene on the mother’s chromosome is erased and then imprinted with a paternal mark such that both gene copies in sperm carry the appropriate paternal mark.
• Mechanism : During its passage from the male and female parents, the autosomal and X chromosomes are stamped differently for different parents i.e. male and female parents so that the chromosomes bear the marking of the parents from which they are originating from. Much of the autosomes from the two parents are similar in genetic constitution but behaviorally they differ in the manner of expression.
• Cause of Genetic imprinting : Methylation
- It results due to direct modification of DNA through methylation. Greater the degree of methylation lesser is the chance of expression of a gene.
- It is generally understood that imprinting affects a chromosome which survives mitosis but not meiosis.
Examples of Genome imprinting
- Huntington chorea : It is an autosomal disease hence the source of this gene for offspring can be either paternal or maternal. In both cases whether paternal or maternal, the symptoms of the disease will be same. But, the time of initiation of symptoms and severity will be different depending upon whether the gene is paternal or maternal.
- Paternal inheritance of a deletion of this region is associated with Prader-Willi syndrome (characterised by hypotonia, obesity, and hypogonadism). of the same deletion is associated with Angelman syndrome (characterised by epilepsy, tremors, and a perpetually smiling facial expression).
Characteristics of Genome imprinting
- (1) Imprinted genome expresses differentially.
- (2) It is erasable i.e. not a permanent modification e.g. gene imprinting in gonadal tissue.
- (3) It is most common in mammals & flowering plants.
- (4) It is species specific.
- (5) Genetic imprinting is not a rule-it is discovered only in some male and female genes. E.g. gene for growth factor is imprinted on mice and not humans.
Hypotheses on the origins of imprinting
- Parental conflict hypothesis: Also known as the kinship theory of genomic imprinting, this hypothesis states that the inequality between parental genomes due to imprinting is a result of the differing interests of each parent in terms of the evolutionary fitness of their genes. The father’s genes that encode for imprinting gain greater fitness through the success of the offspring, at the expense of the mother. The mother’s evolutionary imperative is often to conserve resources for her own survival while providing sufficient nourishment to current and subsequent litters. Accordingly, paternally expressed genes tend to be growth-promoting whereas maternally expressed genes tend to be growth-limiting. In support of this hypothesis, genomic imprinting has been found in all placental mammals, where post-fertilisation offspring resource consumption at the expense of the mother is high.
- Another hypothesis proposed is that some imprinted genes act co-adaptively to improve both fetal development and maternal provisioning for nutrition and care. In it, a subset of paternally expressed genes are co-expressed in both the placenta and the mother’s hypothalamus. This would come about through selective pressure from parent-infant coadaptation to improve infant survival. Paternally expressed 3 (PEG3) is a gene for which this hypothesis may apply.
- Others have approached their study of the origins of genomic imprinting from a different side, arguing that natural selection is operating on the role of epigenetic marks as machinery for homologous chromosome recognition during meiosis, rather than on their role in differential expression. This argument centers on the existence of epigenetic effects on chromosomes that do not directly affect gene expression, but do depend on which parent the chromosome originated from. Natural selection for genomic imprinting requires genetic variation in a population. A hypothesis for the origin of this genetic variation states that the host-defense system responsible for silencing foreign DNA elements, such as genes of viral origin, mistakenly silenced genes whose silencing turned out to be beneficial for the organism. There appears to be an over-representation of retrotransposed genes, that is to say genes that are inserted into the genome by viruses, among imprinted genes. It has also been postulated that if the retrotransposed gene is inserted close to another imprinted gene, it may just acquire this imprint.
Significance of Genome imprinting
- (1) Genome imprinting helps to know the effect of defection in different sexes.
- (2) It shows variations in diseases according to gender.
- (3) To know the source of allele.
Conclusion : Thus genome imprinting helps in understanding the source of allele whether from paternal or maternal source. Moreover chances of survivability depend upon marking pattern of genes. If genes are not imprinted while transmission from one generation to other, their survival capacity declines.
Theories Elaboration: extra information
The kinship theory: The kinship theory is a kin selection model, where matrigenic and patrigenic alleles experience different patterns of relatedness in the social environment (for example, individuals tend to encounter more matrilineal than patrilineal relatives), and as a result, their expression has different consequences for their respective inclusive fitnesses (Haig, 2002, pp 5–6; Figure 1). The kinship theory focuses on genes whose expression level governs the extent of some physiological or behavioral interaction between individuals. As an example, fetal expression of a growth factor not only influences the fetus’ development but may also indirectly affect the growth, and potentially the fitness, of siblings through its demand for shared maternal resources. Variation in the fetal expression level of this growth factor gene can have different, and opposing, inclusive fitness effects for the two alleles if these have unequal relatedness to individuals with whom the fetus interacts (for example, because the fetuses are maternal half-sibs, which share matrigenic but not patrigenic alleles). Because of this relatedness asymmetry, increased expression might increase the inclusive fitness of the patrigenic allele but decrease the inclusive fitness of the matrigenic allele. Haig (1997) termed this selection pressure parental antagonism, drawing the analogy with sexual antagonism, in which a trait is favored in males but disfavored in females (and vice versa). In the face of parental antagonism, an allele that adjusts its expression level in cis to bring about that parental origin’s optimum is selectively favored over an alternative allele that does not adjust its expression (Haig and Westoby, 1989). Assuming parental origin specific expression is possible, the kinship theory then seeks an evolutionarily stable strategy (ESS) for gene expression from the two parental origins (Haig, 1997). This ESS corresponds to an expression level where only the copy from the parental origin that originally favored a higher level of expression is expressed, with the copy from the parental origin that originally favored reduced expression being completely silenced—that is, the canonical pattern of genomic imprinting. Further, the expressing parental origin expresses at that parental origin’s optimal level. At the ESS in this example, the growth factor gene will be expressed exclusively from the patrigenic allele at the patrigene’s optimal level. This sketch of the logic of the kinship theory covers both the strong and the weak versions of the theory (sensu Wilkins and Haig, 2003). These versions correspond to the familiar distinction in evolutionary biology between origin and maintenance: the strong version of the kinship theory claims that the origin of imprinted expression is a consequence of parental antagonism; the weak version submits that once imprinted expression has already been established at a locus, an imprinted gene evolves subject only to the inclusive fitness consequences for the matrigene or patrigene, whichever copy remains expressed. Although Haig and Westoby (1989) originally focused on multiple mating as the source of relatedness asymmetry for matrigenes and patrigenes, later work has extended the kinship theory to cover other sources, including haplodiploidy (Haig, 1992; Queller, 2003) and sex-biased dispersal or sex-biased fitness variance (Haig, 2000b; U´ beda and Gardner, 2010, 2011, 2012; Van Cleve et al., 2010; Brandvain et al., 2011). Further, the kinship theory can be applied to any behavior that influences the fitness of kin and is not limited to the solicitation behaviors of offspring. For example, the provisioning of shared or potentially shared resources by adults is another social behavior that has parentally antagonistic inclusive fitness effects when the two alleles of the provider are asymmetrically related to the recipients of the resource (U´ beda and Gardner, 2011, 2012).
The sexual antagonism theory: The sexual antagonism theory for the evolution of genomic imprinting relies on sex-specific selection pressure (Day and Bonduriansky, 2004; Bonduriansky, 2007; Figure 2). When selection differs between males and females—be it in magnitude or in direction—the allele frequencies in eggs and sperm (and, therefore, in matrigenes an patrigenes of the next generation) become unequal. Two scenarios can generate this non-equivalence. First, if selection is sexually antagonistic for a gene, patrigenes will be enriched for alleles of male benefit and matrigenes will be enriched for alleles of female benefit. There is thus a selective advantage to a novel modifier that adjusts expression level in an imprinted and sex-specific way—that is, patrigenic expression of the gene when in males and matrigenic expression when in females. Second, there is a net selective advantage to modifying expression in the standard imprinted fashion—that is, independently of the sex of bearer—provided that the allele deriving from the parent that experienced stronger selection in the previous generation is expressed. Analogous logic underlies Spencer and Clark’s (2006) ‘Chip Off the Old Block’ theory for the origin of mprinting, where selective asymmetry arises because the haploid genome inherited from the parent of the less dispersive sex will tend to be enriched for locally adapted alleles (see also Spencer and Clark, 2014). Spencer and Clark’s (2006) model therefore predicts the imprinted silencing of alleles inherited from the more dispersive parental sex for loci with key functional roles in the exploitation of the local ecological niche. In the sexual antagonism theory, selection favors imprinted expression because the two alleles at a diploid locus carry, on average, different information or instructions, and one of these copies provides more adaptive information than the other. Population variation for such alleles, and the absence of other epigenetic mechanisms that temper their sex-specific effects on fitness, results in intralocus sexual conflict, whereby a sex is prevented from reaching its phenotypic optimum as a result of a different pattern of selection on the same locus in the other sex (Bonduriansky and Chenoweth, 2009). Whenever there is differential selection between the sexes, the parent of the same sex as the offspring will, on average, provide fitter alleles to that offspring compared with the other parent. Imprinted silencing, therefore, provides a simple mechanism through which individuals can express the more adaptive set of instructions for their sex. For the sexual antagonism theory, imprinting mitigates intralocus sexual conflict by allowing each sex to approach more closely its sex-specific phenotypic optimum. The requirement for the evolution of imprinted expression under this theory is simple: sex-specific selection pressure on a gene. Any species with two sexes is subject to imprinting of genes under sex-specific selection. The requirements of the sexual antagonism theory are less restrictive than any of the other theories’ requirements, and, as such, the sexual antagonism theory is expected to apply more broadly.
The maternal–offspring coadaptation theory: The fitness of a genotype depends not only on its direct phenotypic effect on its bearer but also on the interactions between the phenotypes of different individuals (Moore et al., 1997; Wolf et al., 1999a; McGlothlin et al., 2010). Certain of these interactions are more (or less) than the sum of their parts, producing a form of epistasis (Wolf, 1999). Whenever interactions between traits affect fitness, genetic correlations between the interacting traits are expected to evolve (Sinervo and Svensson, 2002), even when the phenotypes belong to different individuals, as in the case of mothers and offspring (Wolf and Brodie, 1998). Wolf and Hager’s (2006, 2009) maternal–offspring coadaptation theory supposes that the two alleles in an offspring code for nonequivalent fitness interactions with the mother, favoring imprinted expression (Figure 3). They examine two scenarios, one where the same locus pleiotropically influences both maternal and offspring traits (the single-locus model) and a second where different loci affect the maternal and offspring traits (the two-locus model). If the interaction confers higher fitness when there is a greater genetic covariance (which, for the single-locus case, is equivalent to phenotypic ‘matching’) between the two actors, then selection favors expression of the matrigenic allele and silencing of the patrigenic allele in the offspring. In the single-locus model, the two alleles in the offspring at a diploid locus are non-equivalent for fitness because an offspring’s maternally derived allele is necessarily more likely to be present in the mother than its paternally derived allele, and expression of the maternally derived allele is more likely to produce a phenotypic ‘match’ between mother and offspring. In the two-locus model, past correlational selection on the maternal–offspring interaction results in a scenario where the matrigenic portion of the offspring’s genotype is positively genetically correlated with any maternal effects with which the offspring’s phenotype must interact; that is, past selection will have built linkage disequilibrium between the two loci. The interaction is therefore more likely to be a high fitness one when expression is limited to the matrigenic allele, which will have evolved some degree of coadaptation with the alleles present in the mother. If instead the highest fitness interaction is achieved by a more negative genetic covariance (or heightened mismatch) between mother and offspring, then the prediction is for the matrigenic allele to be silenced and the patrigenic allele to be expressed.