- DNA fingerprinting was invented in 1984 by Professor Sir Alec Jeffreys after he realised you could detect variations in human DNA, in the form of these minisatellites.
- DNA fingerprinting is a technique that simultaneously detects lots of minisatellites in the genome to produce a pattern unique to an individual. This is a DNA fingerprint.
- Minisatellites are short sequences (10-60 base pairs long) of repetitive DNA that show greater variation from one person to the next than other parts of the genome. This variation is exhibited in the number of repeated units or ‘stutters’ in the minisatellite sequence. The probability of having two people with the same DNA fingerprint that are not identical twins is very small.
- Just like your actual fingerprint, your DNA fingerprint is something you are born with, it is unique to you.
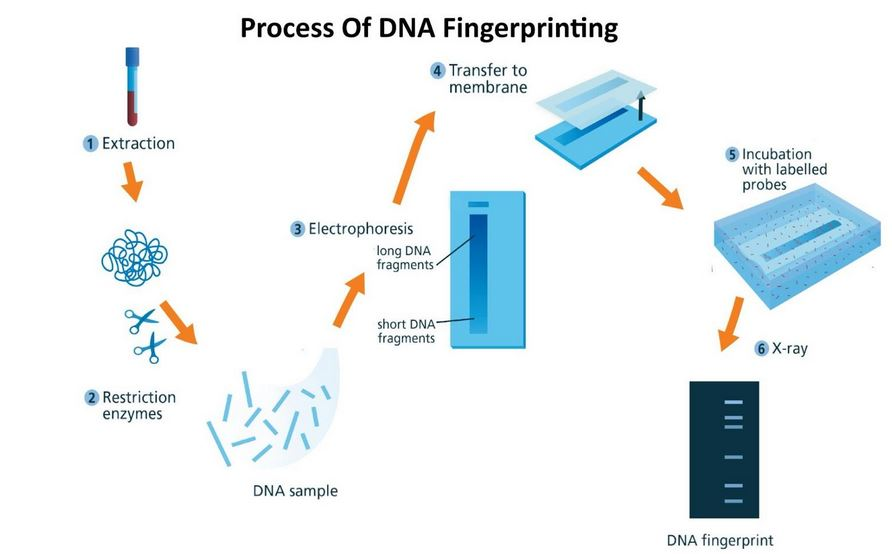
How was the first DNA fingerprint produced?
- The first step of DNA fingerprinting was to extract DNA from a sample of human material, usually blood.
- Molecular ‘scissors’, called restriction enzymes, were used to cut the DNA. This resulted in thousands of pieces of DNA with a variety of different lengths.
- These pieces of DNA were then separated according to size by a process called gel electrophoresis:
- The DNA was loaded into wells at one end of a porous gel, which acted a bit like a sieve.
- An electric current was applied which pulled the negatively-charged DNA through the gel.
- The shorter pieces of DNA moved through the gel easiest and therefore fastest. It is more difficult for the longer pieces of DNA to move through the gel so they travelled slower.
- As a result, by the time the electric current was switched off, the DNA pieces had been separated in order of size. The smallest DNA molecules were furthest away from where the original sample was loaded on to the gel.
- Once the DNA had been sorted, the pieces of DNA were transferred or ‘blotted’ out of the fragile gel on to a robust piece of nylon membrane and then ‘unzipped’ to produce single strands of DNA.
- Next the nylon membrane was incubated with radioactive probes.
- Probes are small fragments of minisatellite DNA tagged with radioactive phosphorous.
- The probes only attach to the pieces of DNA that they are complementary to – in this case they attach to the minisatellites in the genome.
- The minisatellites that the probes have attached to were then visualised by exposing the nylon membrane to X-ray film.
- When exposed to radioactivity a pattern of more than 30 dark bands appeared on the film where the labelled DNA was. This pattern was the DNA fingerprint.
- To compare two or more different DNA fingerprints the different DNA samples were run side-by-side on the same electrophoresis gel.
Uses
Since it was invented in 1984, DNA fingerprinting most often has been used in court cases and legal matters. It can:
- Physically connect a piece of evidence to a person or rule out someone as a suspect.
- Show who your parents, siblings, and other relatives may be.
- Identify a dead body that’s too old or damaged to be recognizable.
DNA fingerprinting is extremely accurate. Most countries now keep DNA records on file in much the same way police keep copies of actual fingerprints. It also has medical uses. It can:
- Match tissues of organ donors with those of people who need transplants.
- Identify diseases that are passed down through your family.
- Help find cures for those diseases, called hereditary conditions.
Fingerprint Test
To get your DNA fingerprint, you would give a sample of cells from your body. This can come from a swab inside your mouth, from your skin, the roots of your hair, or your saliva, sweat, or other body fluids. Blood is usually the easiest way. Lab workers treat the sample with chemicals to separate the DNA, which is then dissolved in water.